15 Apr 2022
Automated assessment of pathology image quality
The new Oxford-developed artificial intelligence tool PathProfiler automates the quality control of large retrospective pathology image datasets to increase their usability in downstream research.
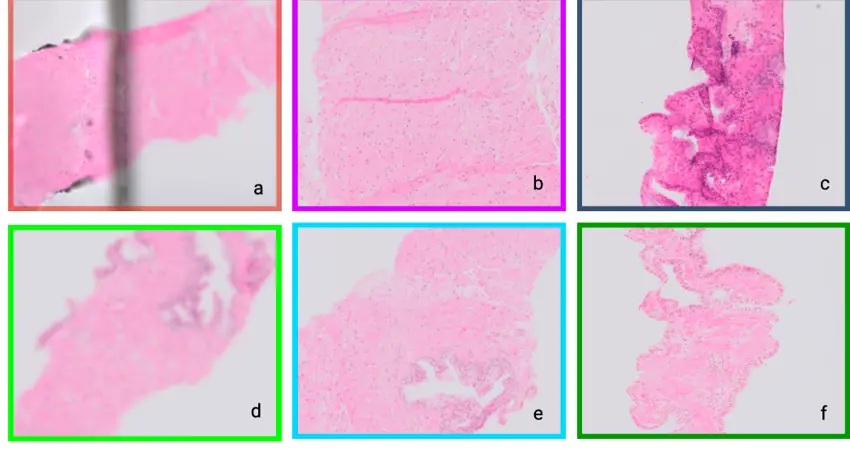
During investigations for cancer, a tissue biopsy may be taken and studied under the microscope to identify abnormal cells. This can help to diagnose and characterise cancer if present, and guide treatment selection.
In more recent times the development of diagnostic digital pathology has gained interest, whereby instead of a light microscope for assessment of a histopathology image, the image is digitised and viewed on a computer screen. Although to date there are limited NHS Cellular Pathology departments with access to such facilities (Oxford being one such centre within the PathLAKE consortium), there is an increasing drive towards the use of digital pathology within the clinical setting, which brings with it the potential to use artificial intelligence (AI) tools.
To realise this opportunity, there has been a surge in research focused on developing and using AI to support pathologists in the assessment of diagnostic biopsies and also in uncovering new disease biology. However, to develop and train AI algorithms requires large datasets of digitised pathology images accompanied by clinical outcome data. Gathering such large datasets requires collating images from multiple diagnostic laboratories, which introduces potential variability in image quality and the risk of a negative impact on downstream analysis. Furthermore, digitised images are often collated retrospectively from stored histopathology glass slides that are subject to age- and storage-related imaging artefacts.
Image quality in the training dataset is known to have an impact on the performance of AI models and the reliability of the generated results. For this reason, it is important to assess image quality during the development and use of AI algorithms. Currently reliant upon manual assessment by pathologists, this is unsustainable given the need for increasingly large datasets; an automated tool is needed to assess image quality that takes into account a combination of potential artefacts and recognises the potential for an intervention to improve the quality.
To address this need, lead author Dr Maryam Haghighat (Institute of Biomedical Engineering, Department of Engineering Science) and Professor Jens Rittscher (Institute of Biomedical Engineering, Big Data Institute and Ludwig Oxford) worked with pathologists Dr Lisa Browning (Department of Cellular Pathology, Oxford University Hospitals (OUH) NHS Trust) and Professor Clare Verrill (Nuffield Department of Surgical Sciences and Department of Cellular Pathology, OUH).
Using retrospective pathology images from the ProMPT (Prostate Cancer Mechanisms of Progression and Treatment) prostate cancer cohort, the team developed PathProfiler, an AI tool to automate quality assessment including a range of potential image artefacts. The tool outputs a usability score, which will help to guide whether an image can be included in the AI training dataset. The scores from the AI algorithm and manual scoring by three expert pathologists were highly correlated (0.89).
To further test PathProfiler, the team assessed the Cancer Genome Atlas prostate and FOCUS colorectal cancer cohorts. In addition to providing a quality score and indicating quality-impacting artefacts that are present, PathProfiler is also able to predict which images could be improved for example, by re-scanning or re-staining. This prediction becomes important particularly for highly curated retrospective cohorts such as those used in prostate cancer research.
The PathProfiler software is available to the public so that other groups can use it for their own research and contribute to the further development of this methodology. The team now plan to further optimise the model, including taking into account additional artefacts observed in other tissue types and cohorts, and to assess the performance and utility of the tool within a clinical pathology digital pipeline.
This study is published in Scientific Reports.
This article is reproduced with kind permission from the Oxford Centre for Early Cancer Detection. The original article is here.




